
Innumerable are the times that the quote “blame it on my genes!” has crossed my mind in response to a sudden pang of envy – be it at the perennially skinny friend whose skeletal frame seemed to betray the tantalizing cheesy wonder of a pizza that they could devour guilt free or be it at the ones so vertically blessed that they have to bend down from realms beyond the Milky Way to have an eye to eye conversation with diminutive me! Though this kind of an arrangement seems to be rather unfair as far as views of equality go, it does seem wise when the purpose of natural selection is considered, that we all have a completely unique genome that may ensure that at least some of our species may survive in the event of a future apocalypse.. It has been said that genes load the gun while our lifestyle pulls the trigger.
Genes in fact are the unknown variable that coupled with lifestyle determinants influence the individual’s response to drugs in ways that can’t be fathomed by mere empirical techniques. Yes indeed therapeutics would be so much easier if a dose of a certain drug produces the expected response in all individuals, however since the reality encompasses a vast variety of factors all influencing the response of an individual to a drug, the next logical step needed is to use this diversity in drug responses to our advantage in providing the best possible patient care and that is exactly what the field of pharmacogenetics aims to achieve.
HISTORICAL PERSPECTIVE
The concept of different drugs producing different responses in different individuals is not new. As early as 510 BC Pythagoras recognized that some individuals developed hemolytic anemia on the consumption of seemingly harmless fava beans.
It however took several hundred years for the reason to be discerned as a deficiency of the enzyme glucose -6 phosphate dehydrogenase in certain individuals that predisposed them to the harmful effects caused by consumption of not just fava beans but a multitude of other drugs such as primaquine. The term pharmacogenetics itself is relatively recent and was first brought into existence by Friedrich Vogel in 1959.
CAUSES FOR VARIATIONS IN DRUG RESPONSE
VARIATIONS IN ENZYMES
Enzymes involved in phase one reactions
It is well known that almost 75% of the drugs that we take are metabolized in phase one bio transformation reactions by CYP450 enzymes and so it comes as no surprise that mutations in the genes encoding these enzymes can lead to transcription of enzyme phenotypes with markedly different activity levels. Mutations can lead to alleles with increased function or decreased function or even duplications of entire genes leading to phenotypes such as PM, IM, EM and UM with enzyme activities increasing from left to right. A highly cited example of such genetic polymorphism would be that of the CYP2D6- A major enzyme that transforms almost one fourth of all known drugs, especially the basic ones like opioids and beta blockers this enzyme ha been touted for the high degree of variability that patients describe on taking a very common antitussive medication- Codeine. CYP2D6 is responsible for the O methylation that converts codeine into its 100 times more potent analgesic morphine. The UM’s and the EM’s respond quickly to the analgesia and experience satisfactory pain relief however that comes at a price of greater side effects like drowsiness and respiratory depression while the PM’s and IM’s may keep on asking for more medication simply because there is not enough morphine formed in their body to alleviate the pain- something which may even make them seem like a drug abuser asking for more drugs in a clinical setting however due to no fault of their own.
Phase 2 enzymes also show polymorphisms. A famous example would be that of UGT1A1- Uridine 5 diphosphoglucuronosyl transferase and TPMT- thiopurine methyltransferase. Both of these enzymes have a role in metabolizing chemotherapeutic drugs used to treat cancer. UGT1A1 metabolites irinotecan a topoisomerase inhibitor and its active metabolite SN38 which limits the dose that can be used due to bone marrow suppression and GI toxicity can present with an extremely poor outcome in PM’s and IM’s who can present with exaggerated neutropenia and diarrhea that can seem unexpected for a certain dose. In addition it also plays a role in physiological conjugation of bilirubin in the liver and a congenital reduction in activity takes the form of the benign disorder known as Gilbert syndrome. On the other hand TPMT exerts its toxic effects through TGN metabolites in response to thiopurine drugs like 6 – mercaptopurine or azathioprine.
This discussion would definitely be incomplete without a mention of some other enzymes that too have high clinical implications in pharmacotherapy. One such is G6PD – the oldest among all enzyme polymorphisms to be noticed by people.
In response to oxidative stressors like fava beans, infection or drugs the levels of this enzyme increase to meet NADPH demands in normal individuals. However in the malaria endemic belt of the world located mainly in parts of Africa and Asia( including parts of India) people seem to have acquired a trait that makes the enzyme less functional and the roc’s more susceptible to hemolysis – most probably as a way of protection against the plasmodium parasite that lives most of its life cycle within the rbcs and hence reducing the lifespan of the rbcs would be a logical way to get rid of the infestation – similar to the reason for high prevalence of sickle cell anemia in these very parts of the world. A rare and intriguing case of how an endemic disease has led to a natural selection of certain genetic polymorphisms that would otherwise not offer a survival advantage and may even be harmful.
Yet another report that is closer to home is the genetic predilection of the Arya Vysya community of southern India to prolonged sedation and even death due to cardiac arrest or apnea on administration of skeletal muscle relaxants like succinylcholine owing to their lower levels of pseudocholineseterase enzyme that metabolizes the drug. This trait has remained in the community due to high degree of intra caste marriages (an artificial selection of a kind).
ALTERATIONS IN PROTEINS
Several proteins too have been implicated as a cause for varied drug responses. OATP1B1 is a transporter located on hepatocytes on the sinusoidal facing membrane which has been found to lead to increased risk for myopathy in response to the use of statins in those with an ineffective transporter. Similarly in a type of AML known as AML M3 or acute promyelocytic leukemia a genotype involving a t(15:17) is associated with the formation of a PML RARa chimeric gene receptor and this responds well to ATRA( all trans retinoids acid) and hence has a better prognosis than another variant called PZFL RARa which is a result of a t(11:17) and has a worse prognosis owing to its lackadaisical response to treatment with ATRA. Such is in fact the case for a majority of cancers and so pharmacogenomics holds great promise as far as development of new cancer chemotherapeutic agents is concerned.
VARIATIONS IN IMMUNE FUNCTION
Yet another interesting set of variations seen involve components of the immune system such as the HLA type of an individual that is also important when considering matching of an organ donor to a recipient is concerned. HLAB polymorphisms can lead to severe hypersensitivity reactions that manifest in the form of a Steven Johnson syndrome or toxic epidermal necrosis especially with the allele HLA-B*57:01. Similarly IFNL3/IL28 gene testing is used to weigh risk benefit ratio when considering whether to treat a patient infected with HCV with ribavirin or not to improve treatment efficacy.
FACTORS AFFECTING THE BIOLOGICAL MILIEU
In some cases the polymorphism may be in an enzyme or protein that affects the biological milieu. For instance variations in the MTHFR(homocysteinemia may be caused) Factor V, prothrombin etc cause the milieu to become pro thrombotic without actually affecting the actions of prothrombotic drugs like estrogens but may modify the risk of a thrombotic event in the presence of such a drug.
POLYGENIC INFLUENCE
Much as we may like to simply put out a list of commonly implicated genes and their various genotypes with the phenotypic influence and hope for it to serve as a comprehensive enough guide for our day to day clinical practice the scenario is far more complex. What I have elucidated above is simply a set of single gene effects or single allele effects in the modification of drug responses but we as doctors are dealing with patients and not single gene systems- patients who are a complex conglomeration of several genes each of which exerts its own unique influence on the way a drug acts in the patients body- some being significant enough to merit clinical notice while others being not very significant. However the bottom line remains that the response output seen is a summation or an integral of all the single gene effect variables that come together to decide how a patient responds to a drug. A relatively simple example of this fact would be how warfarin- an anticoagulant is affected by more than one gene- namely CYP2C9 that reduces its metabolism and further excretion as well as VKORC1 that leads to lesser quantities of the enzyme VKOR that vitamin K acts upon both of which act in an additive manner potentiating the effects of warfarin and causing increased bleeding side effects that are far greater than what is anticipated for a given dose.
EPIGENOMICS
We have discussed fervently about changes in the primary DNA code that leads to varied responses to drugs however there is also another component that exerts its influence discreetly yet majorly. These are various epigenetic modifications such as methylation of DNA or histone modifications which may also combine with polygenic influences to affect responses to drugs and hence is a pretty promising field for future research.
STUDIES OF POLYMORPHISMS
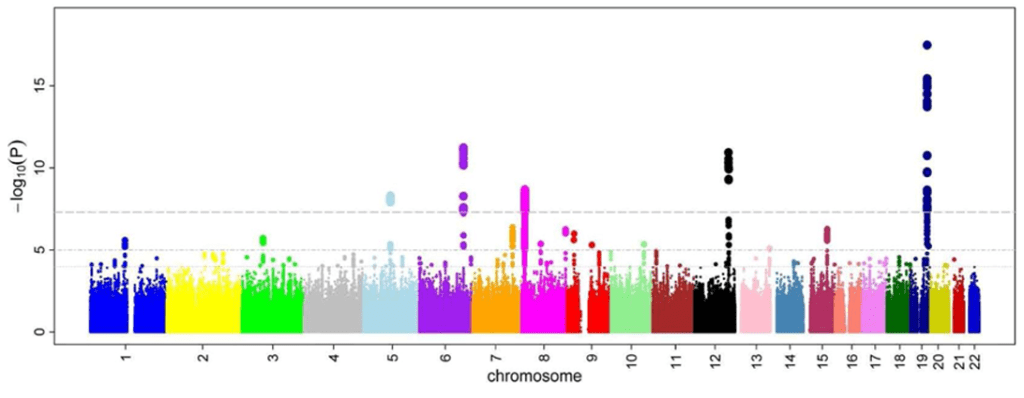
[Manhattan plot – Wikipedia.org]
Now that we have spoken of the various points at which genes exert influence on drug response the next logical step would be to identify the various genetic loci of interest as far as pharmacogenetics is concerned.
There are various methods of doing this. One way is a candidate gene vs a genome wide approach that focuses on what is known regarding the pathways influencing a particular drug and then using a database like PharmGKB to test the hypothesis that variants in a particular set of genes account for varying phenotypes.
This is however time consuming and inefficient because we are only looking for variations in regions of known biological activity and neglecting the rest. Yet another method that is in use is a genome wide association study(GWAS) in which genotypes at almost 500,000 snp sites are compared with a continuous trait or between cases and controls (that is those with and without therapeutic response or an ADR to a drug). An even more comprehensive approach was to couple genomic findings from a GWAS with other large scale measures of gene functions like mRNA levels(transcriptomics) or proteins(proteomics) as the abundance of signals may itself reflect some of the relevant genetic variations.
FUNCTIONAL STUDIES OF POLYMORPHISMS
Once a gene or locus of interest is identified the next step is to discover which coding or regulatory variants contribute. One way of doing this is comparative genomics or actually studying the individual polymorphisms in vitro or in animal models. Highest effects are exerted when a non synonymous SNP or a nonsense or miss ensue mutation leads to a loss of an evolutionarily conserved amino acid residues. With millions of variants being identified from whole genome sequencing studies new innovative methods to identify their functions are also evolving and increasingly computational algorithms have been developed to identify deleterious amino acid substitutions. A real time way of assessing the function of a gene variant would be to artificially construct the variant by site directed mutagenesis and compare the function of the new protein with the normal one.
Yet another set of new technologies that are promising to revolutionize functional gene studies is the ability to form iPSC’s from an individual to study their physiology completely and using the recent Nobel prize winning discovery of CRISPR Cas9 method of gene editing to bring about selected changes in the iPSC’s for the purpose of functional testing.
APPLICATION IN DRUG DEVELOPMENT PROCESS
Majority of clinical trials end up in having to scrap the candidate drug owing to ADR’s that are detected at later stages of the clinical trials leading to immense wastage of resources and time. However if the pharmacogenetic variants that are more likely to respond well to a certain drug candidate can be identified early on in the scheme of events the trials can be tailored to include only that subset of the population and exclude those who receive an unfavourable risk benefit ratio. This can lead to more targeted therapeutic options with better efficacy being delivered to the market and decreased losses for pharmaceutical companies.
Also genomics itself can help in identifying potential targets for drug development process. A worthy example of this would be that when it was found that individuals with lower levels of PCSK9 had lower levels of LDL cholesterol and lesser risk for CAD it led to the evolution of PCSK9 inhibitor antibodies like alirocumab and evolocumab. Similarly were the drugs ivacaftor and lumacaftor developed to target the CFTR in Cystic Fibrosis.
CLINICALLY AVAILABLE PHARMACOGENETIC TESTS
The currently available pharmacogenetic tests aim to identify polymorphisms in HLA types, gene metabolizing enzymes that modify its pharmacokinetics and the gene target proteins. It encompasses a part of what the FDA describes as companion diagnostics – that is a diagnostic test that accompanies a therapeutic drug to determine the applicability of a certain drug to a certain individual.
Currently available tests include abacavir and HLAB57:01, carbamazepine and HLAB15:02, TPMT and thiopurines, CYP2D6 and opioids and tamoxifen, transtuzumab and HER1, imagining, Dasani in and bcr:abl1, warfarin and cyp2c19 and vkorc1 etc.
The clinical pharmacogenomics implementation consortium provides a compiled information database on various drugs and genetic polymorphisms that affect response to them as well as recommendations regarding dosage for each case.
CANCER AND PERSONALISED MEDICINE

In no other field has the advent of personalized medicine being more of a game changer than in oncology. Earlier what was thought of as a single disease entity and given names according to the organ of origin(lung cancer, liver cancer etc) now it has been recognized that cancers are a complex amalgamation of varying genetic defects with such great diversity even in a single tumor in a single individual that they definitely cannot be treated in ‘a one size fits all’ approach. As pathologists come up with newer methods of classifying cancers based on the genetic aberrations responsible for each type pharmacogenetics shall play a great role in the future for achieving a ‘cure for cancer’ which may simply be a more logical and targeted approach at treating each genetic variant specifically rather than a miraculous drug that can claim to cure all kinds of cancer indiscriminately.
Precision medicine targets certain mutations in certain kinds of cancer. For instance the EGFR mutation lung cancer being treated with gefitinib or the ALK positive lung cancer being treated with crizotinib or ceritinib. Transtuzumab being used in HER positive breast cancer and KRAS positive colorectal cancer while Imatinib used for CML. Yet another front runner is immunotherapy for the treatment of cancer which can include checkpoint inhibitors, vaccine immunotherapy and CAR T cell therapy. Checkpoint inhibitors remove the immune systems checkpoints that prevent T cells from attacking cancer cells owing to their mimicking of antigens that are present on normal cells like PDL1. In addition vaccines can be developed to the new antigens expressed on a majority of cancer cells so that the circulating memory cells that react to those antigens are always ready to strike back if the cancer cells begin to show a recurrence thereby acting as sentinels that can eliminate even traces of cancer cells in the body. CAR T is an upgraded version of the same where instead of allowing immunity to develop in Vivo the T cells of the patient are altered ex Vivo to produce chimeric antigen receptors that attach to cancer cell antigens and produce a heightened immune response to them- a technique found effective in many childhood leukemias.
In fact a recent study even found that a universal screening approach for patients rather than a phenotype or clinical symptomatology or family variant based cascade testing approach would lead to higher chances of picking up individuals at risk of cancer however the financial and ethical ramifications of the same are indeed a scope for discussion.
CONCLUSION
Pharmacogenetics opens up its own Pandora’s box of ethical dilemmas regarding the cost vs benefit of genome testing, the fact that such knowledge would compromise insurance cover for people with genetic defects and also the worry that identifying rare diseases with no cures early on would have a negative impact on the mental health of a patient and their loved ones. However it is undisputed that pharmacogenetics is an indispensable part of the future of modern medicine.
We are not really far from an era when each newborn gets his/her their own genome database to be used as a reference for any medication that they may be prescribed throughout their life. An era where personalised prescriptions are the norm rather than the exception, when we doctors wouldn’t just need to hope and pray for a drug to be effective or to not cause an adverse reaction but instead a time when we can actually take logical steps forward in order to ensure the right drugs reach the right patients.
REFERENCES
Jackson SE, Chester JD. Personalised cancer medicine. Int J Cancer. 2015 Jul 15;137(2):262-6. doi: 10.1002/ijc.28940. Epub 2014 May 12. PMID: 24789362.
Pharmacogenomics- Jennifer e hibma and Kathleen m giacommini
Katzung Clinical Pharmacology 14 edition, chapter 5
Principles of gene manipulation and genomics-s.b primrose and r.m twyman, pages 503-506
Rang and Dale’s Pharmacology – 9th edition, pages 152-161
Hans Gerhard Vogel- drug discovery and evaluation : pharmacological assays
3rd edition, volume 1, pages -29-34
Laurence L brunton, randa hilal dandan, Bjorn c knollmann, Goodman and Gillman’s pharmacological basis of therapeutics 13th edition, chapter seven
Manoharan I, Wieseler S, Layer PG, Lockridge O, Boopathy R. Naturally occurring mutation Leu307Pro of human butyrylcholinesterase in the Vysya community of India. Pharmacogenet Genomics. 2006 Jul;16(7):461-8. doi: 10.1097/01.fpc.0000197464.37211.77. PMID: 16788378.
Samadder NJ, Riegert-Johnson D, Boardman L, et al. Comparison of Universal Genetic Testing vs Guideline-Directed Targeted Testing for Patients With Hereditary Cancer Syndrome [published online ahead of print, 2020 Oct 30]. JAMA Oncol. 2020;e206252. doi:10.1001/jamaoncol.2020.6252
Mukherjee S. Genomics-Guided Immunotherapy for Precision Medicine in Cancer. Cancer Biother Radiopharm. 2019;34(8):487-497. doi:10.1089/cbr.2018.2758